The following was originally published in Johns Hopkins Medicine’s Fundamentals.
One thing is certain: COVID-19 isn’t going away any time soon. Although SARS-CoV-2, the virus that causes COVID-19, evolves at a relatively slow pace, infections have been so prevalent around the globe that new strains of the virus have emerged. These new strains may have increased transmissibility, altered virulence or may exhibit resistance to the current vaccines.
Now, scientists are racing to develop better ways to detect emerging SARS-CoV-2 strains among the high number of diagnosed infections.
“Detecting and tracking the genetic changes associated with these new strains is an enormous priority,” says Ben Larman, Ph.D., assistant professor of pathology and director of the Laboratory of Precision Immunology within the Institute for Cell Engineering at the Johns Hopkins University School of Medicine. “Initially, my lab was developing a scalable strain detection technology so that we could discriminate between related viruses. It quickly became clear how important this would be for COVID-19.”
Larman and his team have been working to improve the analysis of genetic material called RNA that forms the genome of many viruses. RNA is not only a component of viruses; it is essential to every form of life on earth.
Specifically, the Johns Hopkins team is developing a technique to scan biological specimens, including saliva or nasal swabs, using specialized DNA probes that sift through a complex “forest” of RNA sequences. The probes can detect specific RNA sequences of viruses and other disease-causing pathogens.
A key feature of the newly-developed test, says Larman, is its ability to analyze and detect the many subtle changes that can occur in the SARS-CoV-2 viral genome — so-called variants, such as those first identified in the United Kingdom and South Africa.
Larman and his team named their test “cRASL-seq” (pronounced krazzle-seek) which stands for capture RNA-mediated oligonucleotide annealing selection and ligation with next generation DNA sequencing.
A report describing the development and application of cRASL-seq, led by Larman and postdoctoral fellow Joel Credle, Ph.D., appeared online Feb. 3 in the journal Modern Pathology. Johns Hopkins Technology Ventures is pursuing patent protection on the technology, which is being licensed to Portal Bioscience LLC, a startup company currently operating in JHTV’s FastForward incubator space.
The cRASL-seq test uses DNA sequencing instruments, which are able to analyze hundreds to thousands of samples at a time. Each test can detect not only SARS-CoV-2, but many other infectious organisms, say the scientists.

Ben Larman
“Sequencing instruments are ubiquitous now,” says Larman. “This type of laboratory test could be adopted by centers all over the world to detect emerging pathogens and even resistance elements associated with bacterial and fungal infections.”
Larman also notes that cRASL-seq skips a step that is required with most of the currently available tests for SARS-CoV-2. Most tests rely on RNA purification kits, which have often been in short supply, hampering efforts to test large swaths of people.
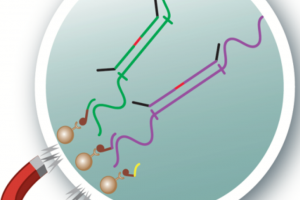
The new test does not rely on such purification kits. Rather, it uses specific probes and magnetic beads to capture target RNA at the same time that the detection probes are binding to the viral RNA.
Looking forward, the Johns Hopkins team is continuing to improve the cRASL-seq technology, expanding the test to detect additional organisms and new SARS-CoV-2 variants as they emerge.
Larman’s laboratory has been part of two teams dedicated to the COVID-19 research effort, which has spanned many departments.
“To see researchers from across the entire university system join forces to combat COVID-19 has been simply awe-inspiring,” says Larman. “It has been a great honor to participate in this response.”